Opening disease X symptom data
(Updates: 2012-04-03, 2012-09-20, 2015-03-14=CC0)
Background
Since my wife started med school (in 1998), I have often questioned the lack of an open, WHO-sponsored database linking symptoms and diseases. As a biologist, it is easy to notice the similarity between diagnostic, multi-entry taxonomic keys (working on characters and taxa), and a key to help diagnose diseases (working with symptoms and diseases). Doctors (or doctors-to-be) have told me reflexively that medicine is an experience-based art, and could never be codified into a database. That a database would ever remove the skill of a doctor has of course never been suggested: just as a taxonomist will use the computer key to create a shortlist of species, to be examined with reference to floras, specimens, and photos, so a doctor would only use such a database as a useful starting point, a preliminary list for a differential diagnosis. Anyone claiming that most doctors can reliably remember a full matrix of thousands of diseases and hundreds of symptoms is being unreasonable. Everyone can use an ‘aide-mémoire’, and there is abundant evidence that checklists in many fields improve efficiency. In countries with only basic medical training, and poor access to medical literature, such a checklist would almost certainly save lives.
Since PalmPilot days, I have been aware of the free Diagnosaurus app, and recent searching has showed me that, of course, this idea has been debated for many years, and has led to a number of implementations, both commercial: DiagnosisPro, Isabel, VisualDX, DXplain, and free (as in beer): DDX-Morph, Diseases Database (see this wikipedia article). However, as far as I am aware, there is still no general open (downloadable, free as in freedom) database for disease-symptom relationships (but see OMIM for heritable diseases). Doesn’t it seem strange that something so fundamentally important should be missing? I am aware of, and accept, the many caveats that need to be applied in building diagnostic systems (well put by the author the Diseases Database), but having open access to these data still seems very important as a global resource for disease-prone humans! In a cynical frame of mind I might deduce that the absence of this database is due to the general trend of the medical industry to close and commercialize anything related to healthcare, but at the same time, I don’t think it is likely that many of the commercial tools listed above are making vast amounts of money. More likely it is due to the same reactionary aspect of traditional medical education that makes ‘evidence-based medicine’ such a revolutionary concept!
Anyway, the bottom line is that a number of doctors who teach at the successful Alam Sehat Lestari (ASRI) clinic in Indonesian Borneo think that a locally-generated disease X symptom database would be a very useful tool for the trainee doctors to have access to, and senior volunteer doctors have also stated interest in populating such a database. So after over ten years of grumbling, I think I might try to implement such a project, humbly accepting that I’m not trained as a physician!
Implementation
So, how to do it right... not just for our own internal needs, but as a basis for, idealistically, a much larger project of sharing these data? As a fan of the Linked Data movement, I immediately wonder how best to make this database truly open and inter-linkable? Key attributes of a strong system include:
- A well-designed, and ideally shared, list of diseases
- A well-designed, and ideally shared, list of symptoms
- Decentralized data acquisition and hosting, combined with the ability to vet data sources
- As simple as possible!
We already use the comprehensive WHO ICD10 list in our patient database, but even more useful there is already an OBO ontology for diseases (DO), which links to ICD and to Dbpedia/wikipedia pages. Similarly, there is an OBO symptom ontology (SYMP). Diseases and symptoms thus have URIs and much of the setup work is therefore done!
There are a few suggestions for linkage properties on the symptom
ontology wiki (e.g., Disease hasClinicalFinding Symptom, Symptom
symptom_of Disease), but I think we need an intermediate Class to
hang optional metadata on, such as:
- Who created the disease X symptom record. This will allow users to assess their confidence in the data.
- When the record was created.
- An estimate of the frequency of either the number of cases of a
disease in which a symptom was observed (
estProbOfSymptom), or the number of cases with an observed symptom in which a disease was found (estProbOfDisease). - The population the record is drawn from. This would optimally point to another ontology for ethnic/geographic populations.
- For internal use: a patient ID/URI and date that the symptom/disease was observed. Records with these data should probably not have estimates of frequency.
- Other notes.
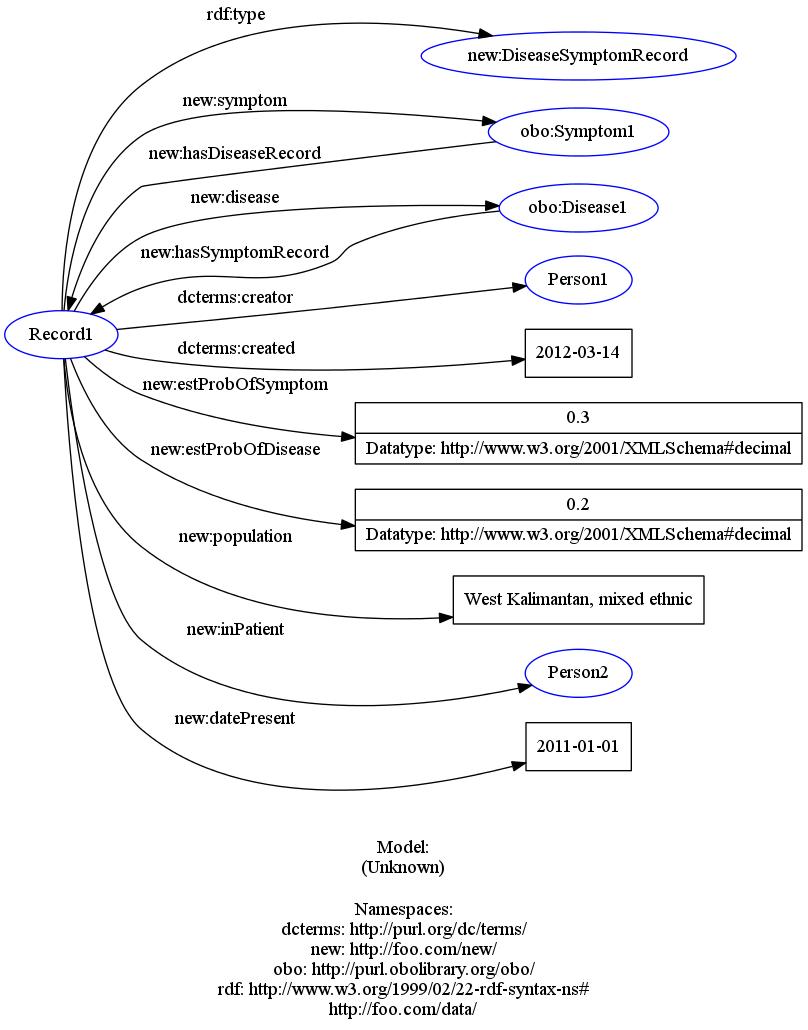
Thus, I propose a basic structure of this nature (here in RDF/Turtle):
@prefix obo: <http://purl.obolibrary.org/obo/> .
@prefix dcterms: <http://purl.org/dc/terms/> .
@prefix new: <http://foo.com/new/> .
@prefix : <http://foo.com/data/> .
:Record1 a new:DiseaseSymptomRecord ;
new:symptom obo:Symptom1 ;
new:disease obo:Disease1 ;
dcterms:creator :Person1 ;
dcterms:created "2012-03-14" ;
new:reference :Document1 ; # Added 2012-04-03
new:estProbOfSymptom 0.3 ;
new:estProbOfDisease 0.2 ;
new:population "West Kalimantan, mixed ethnic" ;
new:inPatient :Person2 ;
new:datePresent "2011-01-01" .
# reverse links:
obo:Disease1 new:hasSymptomRecord :Record1 .
obo:Symptom1 new:hasDiseaseRecord :Record1 .
Deployment
The workflow would thus be:
- Set up local database (probably relational, mySQL) that includes tables for DO and SYMP entries.
- Get the data creators to add records to a
DiseaseSymptomRecordtable, with metadata. - Create lookup applications to use data locally.
- Re-serialize data as RDF (care needed not to include private patient data).
- Either dynamically host local URIs or periodically dump data as a static RDF file.
- Register the local URIs/dump file globally (e.g., Sindice), and with a central project datastore/query platform.
- Query multiple datasets to build a lookup app for doctors,
optionally filtering by data source URI (graph URI) and by
dcterms:creator.
The best future for such a project would have hundreds of data suppliers using the same terms and ontologies (same RDF ‘schema’), with one or more nicely designed database and query engine that harvests all the provider records, filters as required, and offers a nice interface to the doctors. What a useful resource this would be!
Partners
I’ve posted this as an open call for comments/criticism and hopefully to find interested partners. Please post comments here, or send via email.
UPDATE: Beyond the academe (2012-04-03)
I’ve been encouraged by responses to this post to dream a bit more, particularly about ‘filling the matrix,’ the laborious task of assembling the data. What if this project could be sold in a way that it really catches on? The building of a global resource for health could become a global community project, á la Wikipedia. One of the most active communities on the the web are sufferers of diseases, organized into thousands of support groups. If each support group web page included a link to a well-designed public ‘disease X symptom datum’ entry tool, useful information could be collected for even the rarest diseases. While many symptoms are too technical for the patient to record, many are not. Many of the signs and symptoms discoverable in a physical examination could be recognized by a ‘citizen physician’ with careful documentation. And these will be the symptoms most useful for subsequent diagnosis in low-budget medical situations.
This could become the largest citizen science project ever! And as with all citizen science, the data would be... what the data would be! Not valueless, not 100% trustworthy, but somewhere in between, needing careful analysis, and caveat-filled use. Much could be learned from projects like eBird. Potentially the sheer volume of data could overwhelm the danger of uninformed or malicious errors. Additionally, the transcription of symptom lists from the medical literature could be ‘farmed out’ to volunteers. The attributable, distributed nature of the data would always mean that users could include only those sources that they trust.
On the users’ side, in full awareness of the potential problems of presenting the public with complex, unfiltered data, I think direct application of these data could lead to major health and welfare benefits. Both my physician wife and I have been deeply impressed with the work of David Werner and his Hesperian foundation (David actually visited the ASRI clinic in 2007). His book, ‘Where There is No Doctor’ (free, here) may be the single most important, life-saving book in the world, and has definitely helped us with some of our own health issues. While it may be scorned by some professionals, one of the most important things the book does is help people determine when it is vital to seek medical help. It educates, and helps take some fear out of the unknown of an illness. We observe villagers wasting large amounts of their small incomes on unneeded medicine and expensive trips to the local clinics. Imagine a free, well-engineered e-resource, a ‘Where There is No Doctor,’ for the 21st century, that helps diagnose and educate people all over the world. An open diagnosis X disease database would surely underlie such a resource.
UPDATE: The OpenDDx project (2012-09-20)
Please see community progress on this issue at oddx.i2i2t.net.
UPDATE: Public Domain (2015-03-14)
I dedicate all ideas in this particular blog post that might be fairly considered ‘mine’ to the public domain, under CC0. See this post for more.