|
shiba
|
SHIBA main header More...
#include <stdio.h>#include <stdlib.h>#include <unistd.h>#include <ctype.h>#include <math.h>#include <pthread.h>#include <string.h>#include <time.h>#include "version.h"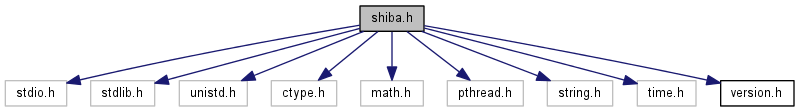
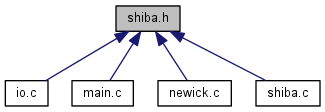
Go to the source code of this file.
Data Structures | |
| struct | config |
| The structure that stores the configuration information. More... | |
| struct | phylo |
| A general phylogeny structure returned from pasring Newick. More... | |
Macros | |
| #define | _GNU_SOURCE |
| to allow the very useful asprintf() | |
| #define | Sasprintf(write_to,...) |
| Safer asprintf macro for extending strings. | |
Functions | |
| void | readArgs (int argc, char **argv) |
| Parses arguments to the executable. | |
| void | readXML () |
| Open file and parse XML. | |
| void | printIndata () |
| Output matrices for checking by user. | |
| phylo | parseNewick (char *in) |
| void | phyloToLineage (phylo p) |
| Reconciles the phylogeny edges with the geo time periods | |
| void | help () |
| Default help output. | |
| void | error (char *a) |
| Basic error output and exit function. | |
| int * | mem1d_i (int dimx) |
| Dimensions an initialized 1-D array of int. | |
| double * | mem1d_d (int dim) |
| Dimensions an initialized 1-D array of doubles. | |
| int ** | mem2d_i (int dimx, int dimy) |
| Dimensions an initialized 2-D array of int. | |
| void | free2d_i (int **ptr, int dimx) |
| int ** | mem2d1_i (int dimx) |
| Dimensions a initialized 1-D array of int pointers, ready to be allocated using mem1d_i() . | |
| void | free2d1_i (int **ptr, int dimx) |
| Frees the ragged array allocated by mem2d1_i(). | |
| double ** | mem2d_d (int dimx, int dimy) |
| Dimensions an initialized 2-D array of doubles. | |
| void | free2d_d (double **ptr, int dimx) |
| char ** | mem2d1_c (int dimx) |
| Dimensions a initialized 1-D array of character pointers, ready to be allocated using asprintf. | |
| void | free2d1_c (char **ptr, int dimx) |
| Frees the ragged array allocated by mem2d1_c(). | |
| double *** | mem3d_d (int dimx, int dimy, int dimz) |
| Dimensions an initialized 3-D array of doubles. | |
| void | free3d_d (double ***ptr, int dimx, int dimy) |
| int *** | mem3d_i (int dimx, int dimy, int dimz) |
| Dimensions an initialized 3-D array of int. | |
| void | free3d_i (int ***ptr, int dimx, int dimy) |
| void | shiba (phylo p) |
| double | findMaxArea () |
| double | findMaxDist () |
| void * | biogeo () |
| void | printSuccessAll (phylo p) |
| double | pDispersal (double x) |
| double | pSurvival (double x) |
| void | printArray (int t) |
Variables | |
| config | Cfg |
| The main config structure. | |
| int | Times |
| The number of time periods. | |
| double * | RealTime |
| The age of the start of the period. | |
| int | Spaces |
| The number of spatial units. | |
| char ** | SpaceName |
| The names of the spatial area. | |
| int | Phylos |
| The number of phylogenies. | |
| char ** | Phylo |
| The phylogeny strings. | |
| int | Taxa |
| The number of terminal taxa. | |
| char ** | Taxon |
| Terminal taxa names. | |
| double ** | Area |
| The areas of spates at each time: Area[Time][Space]. | |
| double *** | Dist |
| The distances among spaces at each time. | |
| double ** | PSurv |
| The prob of survival PSurv[Time][Space]. | |
| double *** | PDisp |
| The prob of dispersal PDisp[Time][Space]. | |
| int *** | Extant |
| 0/1 indcating the existance of a taxon or fossil during a particular time period. Care is needed in assigning fossils to periods, due to reconciliation stage. [x][t][s] | |
| int | Lineages |
| The number of lineages = phylo.nnodes. | |
| int ** | LineagePeriod |
| The 0/1 existence of a lineage in a time period. | |
| int ** | LineageDaughters |
| The daughters of each lineage. | |
| int * | LineageDaughtersN |
| The number of daughters of each lineage. | |
| int *** | LineageExtant |
| Whether each lineage is extant. | |
| int | LinExtantN |
| The number of lineages extant at present. | |
| char * | DataFile |
| Name of the data file. | |
| int | PhyloToUse |
| Index number of the phylogeny currently in use. | |
| int | PrintData |
| Switch 0/1 to control output of raw data. | |
| int ** | record |
| long int | success |
| long int | topresent |
| double | maxdist |
| double | maxarea |
| pthread_mutex_t | mymutex |
| int | RUN_BATCH |
| the number of threads to run in parallel | |
SHIBA main header
Definition in file shiba.h.
| #define Sasprintf | ( | write_to, | |
| ... | |||
| ) |
Safer asprintf macro for extending strings.
From 21st Century C.
Definition at line 13 of file shiba.h.
Referenced by parseNewick(), and readXML().
| void error | ( | char * | error_msg | ) |
Basic error output and exit function.
Note that no freeing is done, possibly leading to memory leaks.
| error_msg | the error message |
Definition at line 612 of file io.c.
Referenced by main(), mem1d_d(), mem1d_i(), mem2d1_c(), mem2d1_i(), mem2d_d(), mem2d_i(), mem3d_d(), mem3d_i(), parseNewick(), phyloToLineage(), readArgs(), and readXML().
| void free2d1_c | ( | char ** | ptr, |
| int | dimx | ||
| ) |
Frees the ragged array allocated by mem2d1_c().
| dimx | The dimensions of the vector |
| void free2d1_i | ( | int ** | ptr, |
| int | dimx | ||
| ) |
Frees the ragged array allocated by mem2d1_i().
| dimx | The dimensions of the vector |
Definition at line 670 of file io.c.
Referenced by main().
| double* mem1d_d | ( | int | dimx | ) |
Dimensions an initialized 1-D array of doubles.
To free, just use free().
| dimx | The dimensions of the vector |
Definition at line 628 of file io.c.
References error().
Referenced by parseNewick(), and readXML().
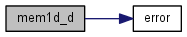
| int* mem1d_i | ( | int | dimx | ) |
Dimensions an initialized 1-D array of int.
To free, just use free().
| dimx | The dimensions of the vector |
Definition at line 643 of file io.c.
References error().
Referenced by parseNewick(), phyloToLineage(), and readXML().
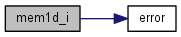
| char** mem2d1_c | ( | int | dimx | ) |
Dimensions a initialized 1-D array of character pointers, ready to be allocated using asprintf.
The result will be a `ragged' matrix. To free, use free2d1_c() .
| dimx | The dimensions of the vector |
Definition at line 685 of file io.c.
References error().
Referenced by parseNewick(), and readXML().
| int** mem2d1_i | ( | int | dimx | ) |
Dimensions a initialized 1-D array of int pointers, ready to be allocated using mem1d_i() .
The result will be a `ragged' matrix. To free, use free2d1_i() .
| dimx | The dimensions of the vector |
Definition at line 658 of file io.c.
References error().
Referenced by phyloToLineage().
| double** mem2d_d | ( | int | dimx, |
| int | dimy | ||
| ) |
| int** mem2d_i | ( | int | dimx, |
| int | dimy | ||
| ) |
Dimensions an initialized 2-D array of int.
Free with free2d_i()
Definition at line 712 of file io.c.
References error().
Referenced by phyloToLineage().
| double*** mem3d_d | ( | int | dimx, |
| int | dimy, | ||
| int | dimz | ||
| ) |
| int*** mem3d_i | ( | int | dimx, |
| int | dimy, | ||
| int | dimz | ||
| ) |
Dimensions an initialized 3-D array of int.
Free with free3d_i()
Definition at line 793 of file io.c.
References error().
Referenced by phyloToLineage(), and readXML().
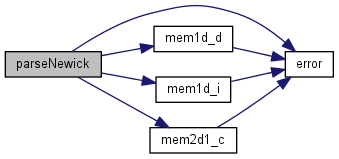
| phylo parseNewick | ( | char * | in | ) |
Definition at line 7 of file newick.c.
References phylo::age, phylo::bl, phylo::depth, error(), mem1d_d(), mem1d_i(), mem2d1_c(), phylo::ndaughter, phylo::nnodes, phylo::parent, Sasprintf, and phylo::taxon.
Referenced by main().
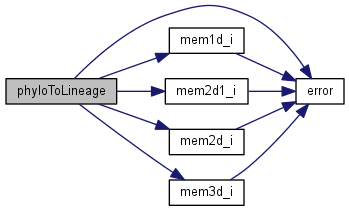
| void phyloToLineage | ( | phylo | p | ) |
Reconciles the phylogeny edges with the geo time periods
For example, imagine three past time periods (A, B, C) and the future (Z), and several phylogeny edges (a-k):
Z : a b d e g h j ..:...|.....|...|.....|...|.....|....|..... NOW : | | | | +--+--+ | C : +--+--+ | | |i | : |c | | +----+--+ ..:......|......|.....|...........|........ : | +--+--+ |k B : | |f | ..:......|.........|..............|........ A : | | |
These would be encoded with reference to the time periods as:
a b c d e f g h i j k
----- ----- ---------
Z | 1 1 0 1 1 0 1 1 x 1 0
C | 0 0 1 1 1 0 0 0 x 0 1
B | 0 0 1 0 0 1 0 0 x 0 1
A | 0 0 1 0 0 1 0 0 x 0 1
Note that we need to push the branching events `forward,' so that there is only ever one root stem passing through the beginning of the first time period.
Definition at line 199 of file newick.c.
References phylo::age, phylo::bl, error(), Extant, LineageDaughters, LineageDaughtersN, LineageExtant, LineagePeriod, Lineages, mem1d_i(), mem2d1_i(), mem2d_i(), mem3d_i(), phylo::ndaughter, phylo::nnodes, phylo::parent, RealTime, Spaces, Taxa, phylo::taxon, Taxon, and Times.
Referenced by main().
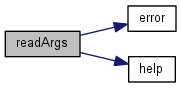
| void readArgs | ( | int | argc, |
| char ** | argv | ||
| ) |
Parses arguments to the executable.
Pass argc and argv directly from main() . copied from the GNU libc getopt() example.
| argc | The number of arguments |
| argv | The array of strings of the various arguments |
Definition at line 69 of file main.c.
References Cfg, DataFile, error(), help(), PhyloToUse, PrintData, and RUN_BATCH.
Referenced by main().
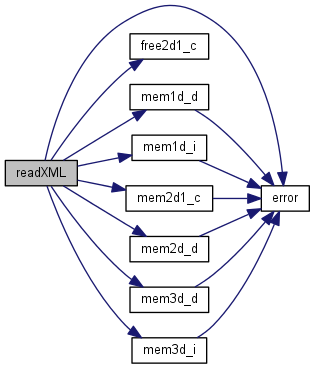
| void readXML | ( | ) |
Open file and parse XML.
Derived from mini-xml website examples.
Definition at line 13 of file io.c.
References Area, Cfg, DataFile, Dist, error(), Extant, free2d1_c(), mem1d_d(), mem1d_i(), mem2d1_c(), mem2d_d(), mem3d_d(), mem3d_i(), Phylo, Phylos, RealTime, RUN_BATCH, Sasprintf, SpaceName, Spaces, config::startSpace, Taxa, Taxon, and Times.
Referenced by main().
| char* DataFile |
Name of the data file.
Default: shibaInput.xml
Definition at line 91 of file shiba.h.
Referenced by main(), printIndata(), readArgs(), and readXML().
| int** LineageDaughters |
The daughters of each lineage.
Definition at line 86 of file shiba.h.
Referenced by main(), phyloToLineage(), and printIndata().
| int* LineageDaughtersN |
The number of daughters of each lineage.
Definition at line 87 of file shiba.h.
Referenced by main(), phyloToLineage(), and printIndata().
| int*** LineageExtant |
Whether each lineage is extant.
[l][t][s]
Definition at line 88 of file shiba.h.
Referenced by main(), phyloToLineage(), and printIndata().
| int** LineagePeriod |
The 0/1 existence of a lineage in a time period.
Definition at line 85 of file shiba.h.
Referenced by main(), phyloToLineage(), and printIndata().
| int PhyloToUse |
Index number of the phylogeny currently in use.
Definition at line 92 of file shiba.h.
Referenced by main(), and readArgs().
| int PrintData |
Switch 0/1 to control output of raw data.
Definition at line 93 of file shiba.h.
Referenced by main(), and readArgs().
 1.8.2
1.8.2